|
2/19/2023 0 Comments Drag data points graph igor pro When fluorescently tagged proteins are bleached in a region of interest (ROI), the recovery in fluorescence occurs due to the movement of unbleached SEP-tagged protein from areas outside the ROI into the bleached region. Fluorescence recovery after photobleach (FRAP) relies on the high mobility of cellular proteins under physiological conditions within the plasma membrane proteins undergo various types of motion from free diffusion to flow motion and/or anchoring (see Jaskolski and Henley, 2009). As SEP only absorbs photons when exposed to neutral pH, it allows the selective bleaching of only SEP-tagged plasma membrane proteins ( Ashby et al., 2004a, 2006 Bouschet et al., 2005 Jaskolski et al., 2009 Martin et al., 2008). Like other fluorescent dyes, when illuminated at high intensity, GFP derivatives can be irreversibly bleached without detectably damaging intracellular structures ( Patterson et al., 1997 Swaminathan et al., 1997 Wiedenmann et al., 2009). Fortunately, our experience in neurons is that recombinantly expressed SEP-tagged surface membrane proteins produce bright signals with a very good signal to noise ratio, meaning that surface expressed fluorophore can be readily distinguished ( Ashby et al., 2004a, 2006 Bouschet et al., 2005 Jaskolski et al., 2009 Martin et al., 2008). This is an important issue to be aware of and control for with acid and ammonium chloride wash protocols (see Section 5) that specifically identify the contributions of surface expressed and intracellular proteins. Thus, if only ~5% of the SEP is surface expressed and exposed to pH 7.4 and 95% is intracellular and exposed to pH 5.5, there will be the same levels of fluorescence signal from the cell surface and inside the cell. Quantification of the pH-dependence of SEP fluorescence suggests that it is 21× brighter at pH 7.4 than at 5.5 ( Sankaranarayanan et al., 2000). This is useful because most stages of the secretory pathway in neurons and other cells occur in acidic compartments, so a SEP-tagged membrane protein absorbs and emits fluorescence only when inserted in the plasma membrane ( Ashby et al., 2004b). This allows the selective imaging of tagged proteins exposed to neutral pH environment. 
Protonation of SEP decreases photon absorption and therefore eliminates fluorescence emission at low pH. (2000).Īmong these, superecliptic pHluorin (SEP), a pH-sensitive derivative of eGFP, has been extensively used for live cell imaging of plasma membrane proteins ( Ashby et al., 2004b Sankaranarayanan et al., 2000). (1998) Pakhomov and Martynov (2008) Sankaranarayanan et al. 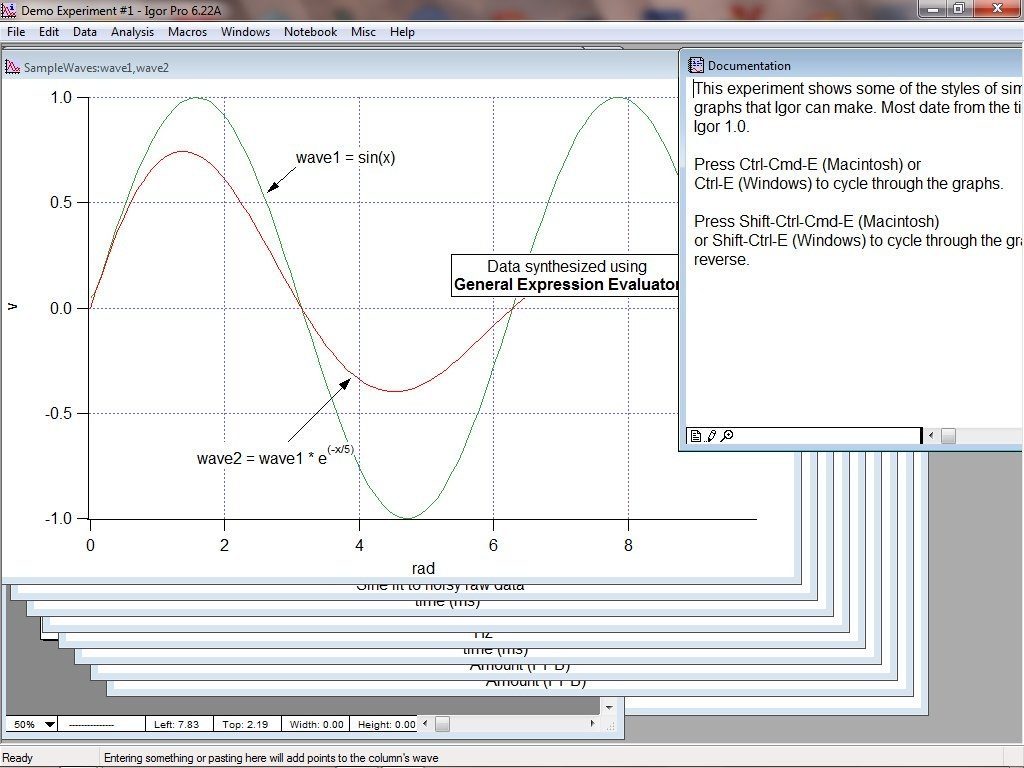
To circumvent these complications, GFP family proteins have been engineered to generate a range of mutants with specifically altered properties such as shifts in emission wavelength or pH sensitivity ( Table 6.1).ĭata taken from Ashby et al. Therefore, an important experimental consideration is that the compartmental localization of the fluorescent signal corresponds to the main site of residence of the mature protein. Thus, fluorescent molecules can be found at all stages of the protein production pathway from early after synthesis until degradation. A potential limitation of most GFP-derived or -related fluorophores, however, is that they are fluorescent as soon as the protein is folded and remain fluorescent until degraded. Crucially, virtually any protein can be tagged with GFP resulting in a fusion that usually retains the same targeting and functional properties as the parent protein when expressed in cells. Ramp Waves.Green fluorescent protein (GFP) and subsequent derivatives and alternatives ( Pakhomov and Martynov, 2008 Tsien, 1998) are bright, stable, nontoxic fluorophores that allow prolonged imaging of protein trafficking in live cells. 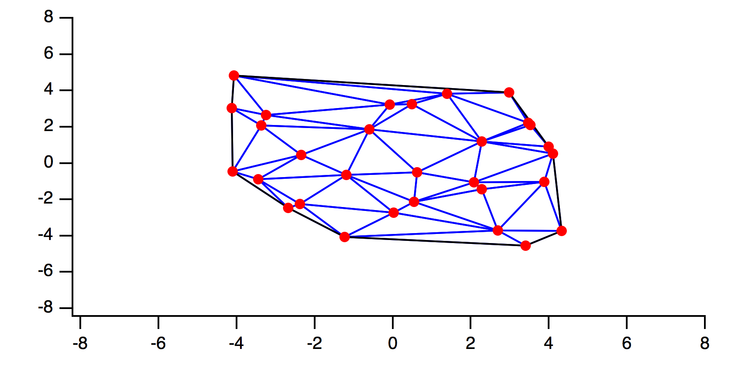
99Ĭhapter 5 Set up Triggering.104ĭigital Triggers.104Īnalog Triggers. PART B-ACQUISITION FUNCTIONS: THE MP MENU. Human Anatomy & Physiology Society Position Statement on Animal Use. What’s new for AcqKnowledge 3.9 for Mac OS™ X. Visit the online support center at TABLE OF CONTENTS 
Data Acquisition and Analysis with BIOPAC MP Systems 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
